AggV3 samples¶
AggV3 was created using single-sample gVCFs generated as part of the DRAGEN 3.7.8 Realignment Project.
The dataset includes 138,399 samples from:
- 100,000 Genomes Project release 19 (including those originally aligned to GRCh37)
- NHS GMS release 4
- COVID release 7
Additionally, seven genomes from Genome in a Bottle (GIAB) have been included for quality validation purposes.
| GIAB sample ID | Library prep | Sex | Person |
|---|---|---|---|
| HG001 | Precision FDA | Female | Utah/Mormon, aka NA12878 |
| HG002 | Precision FDA | Male | Son in Ashkenazim Jewish Trio |
| HG003 | Precision FDA | Male | Father in Ashkenazim Jewish Trio |
| HG004 | Precision FDA | Female | Mother in Ashkenazim Jewish Trio |
| HG005 | Precision FDA | Male | Son in Chinese Trio |
| HG006 | Precision FDA | Male | Father in Chinese Trio |
| HG007 | Precision FDA | Female | Mother in Chinese Trio |
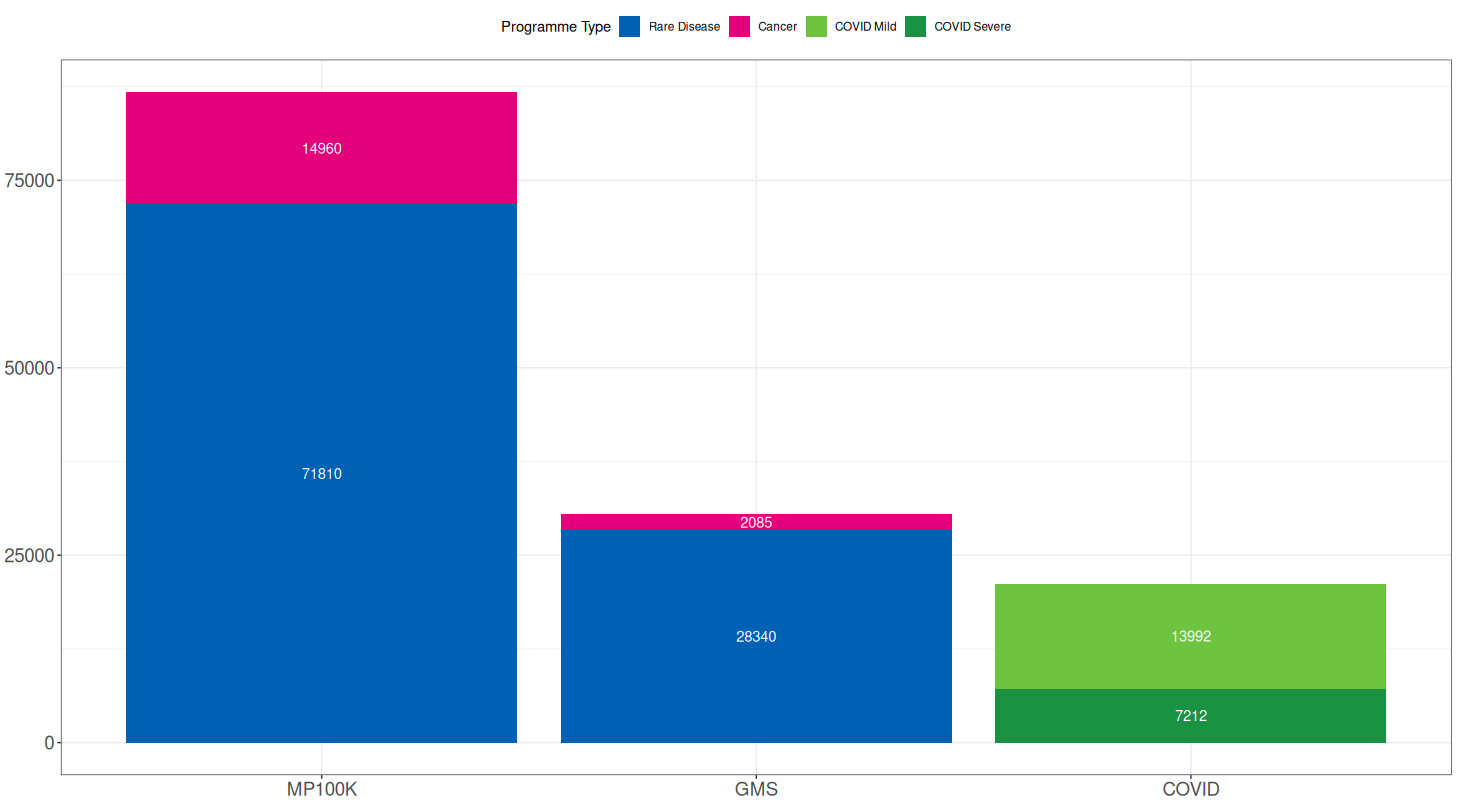
Below is a breakdown of the sample counts from each programme. For the 100,000 Genomes Project and NHS GMS, counts are further split into Rare Disease and Cancer, and for COVID, into Severe and Mild cases:
Participant selection criteria¶
To be included in aggV3, participants had to meet the following criteria:
- they were consented for research in the relevant data release freezes at the time of selection
- they had a successfully aligned germline genome available as part of the DRAGEN 3.7.8 realignment project
As aggV3 integrates data across multiple programmes, some individuals have more than one available germline genome. This can be due to them being enrolled in multiple programmes or, in some cases, having multiple genomes within a single programme. The latter case is common for participants who initially enrolled as part of a rare disease family and were later diagnosed with cancer, resulting in their enrolment under the cancer programme.
We applied the deduplication procedure described below to reduce the likelihood of including the same individual more than once in the aggregate. Despite these efforts, a small number of duplicate samples remain - approximately 15 individuals are duplicated.
Deduplication logic:
- select the most recently delivered genome within each programme
- for individuals enrolled in multiple programmes, select the genome from the programme with the highest priority. The programme types were ranked in the following order:
- NHS-GMS Rare Disease
- 100kGP Rare Disease
- NHS-GMS Cancer
- 100kGP Cancer
- COVID
Associated sample list tables¶
We provide three sample list files associated with aggV3:
- A table listing all participants included in aggV3:
s3://512426816668-gel-data-resources/dragen3.7.8/AggV3_resources/samples/sample_list/2026-01-23/sample_list_aggv3.csv - A table listing the set of individuals considered for inclusion in aggV3 prior to deduplication:
s3://512426816668-gel-data-resources/dragen3.7.8/AggV3_resources/samples/sample_list/2026-01-23/sample_list_considered_for_aggv3.csv - A list of samples consented for research:
s3://512426816668-gel-data-resources/dragen3.7.8/AggV3_resources/samples/consented_individuals/2026-01-23/aggv3_consented_samples.txt
Below we describe the content of the provided sample list tables:
| Column | Description |
|---|---|
participant_id |
Genomics England participant identifier. |
associated_case_reference |
Referral ID/s or "gel_case_reference/s" associated with the participant's GMS case(s). Referral IDs are comma-separated where appropriate. When filtering for a specific ID, ensure that your approach accounts for IDs embedded within a comma-separated string rather than assuming a single value field. |
platekey |
Genomic sample identifier (LP-xxx-DNA_xxx). |
dragen_delivery_id |
Associated dragen_delivery_id that the sample relates to. If NA, sample was not under consideration for realignment. |
dragen_karyotypic_sex_estimation |
Sex karyotype ploidy estimates as provided by DRAGEN 3.7.8 output. If NA, data was not present in the *_ploidy_estimation_metrics.csv or not retrieved. |
type |
experimental/rare disease/cancer/covid mild/covid severe + germline. |
study_source |
GMS, MP100K, COVID. |
programme |
cancer, covid, rare_disease. |
sample_source |
Sample source material (Blood, saliva, etc.). |
sample_preparation_method |
Sample preparation method (EDTA, FF, ORAGENE). |
sample_library_method |
Sequence library preparation method. |
aggv3_inclusion |
TRUE/FALSE. For the AggV3 specific table, these will all be listed as TRUE. |
participant_grouping |
Identifier to group participants which are linked to the same NHS ID and was used by the participant deduplication process. |
multi_programme_ppt_ids |
All participant_id/s associated with the participant_grouping identifier. |
family_grouping |
Extended family group identifier. Example: In GMS if a family consists of two parents + two probands, these would be split in 2 trio-referrals. Referral A has parents + proband_A, and referral B has parents + proband_B. These two distinct referrals would be part of a single grouped family. |
duplicate_of |
We have 18 duplicates (n = 36). This column provides the duplicated platekey of the duplicate pairing. If row is NA, no known duplicate has been identified at this stage. n=6 cases where unique platekeys map to two unique participant IDs, but are the same individual after NHS-ID mapping (MP100K). n=30 cases where unique platekeys match a single unique participant ID (GMS). |
Sample characteristics¶
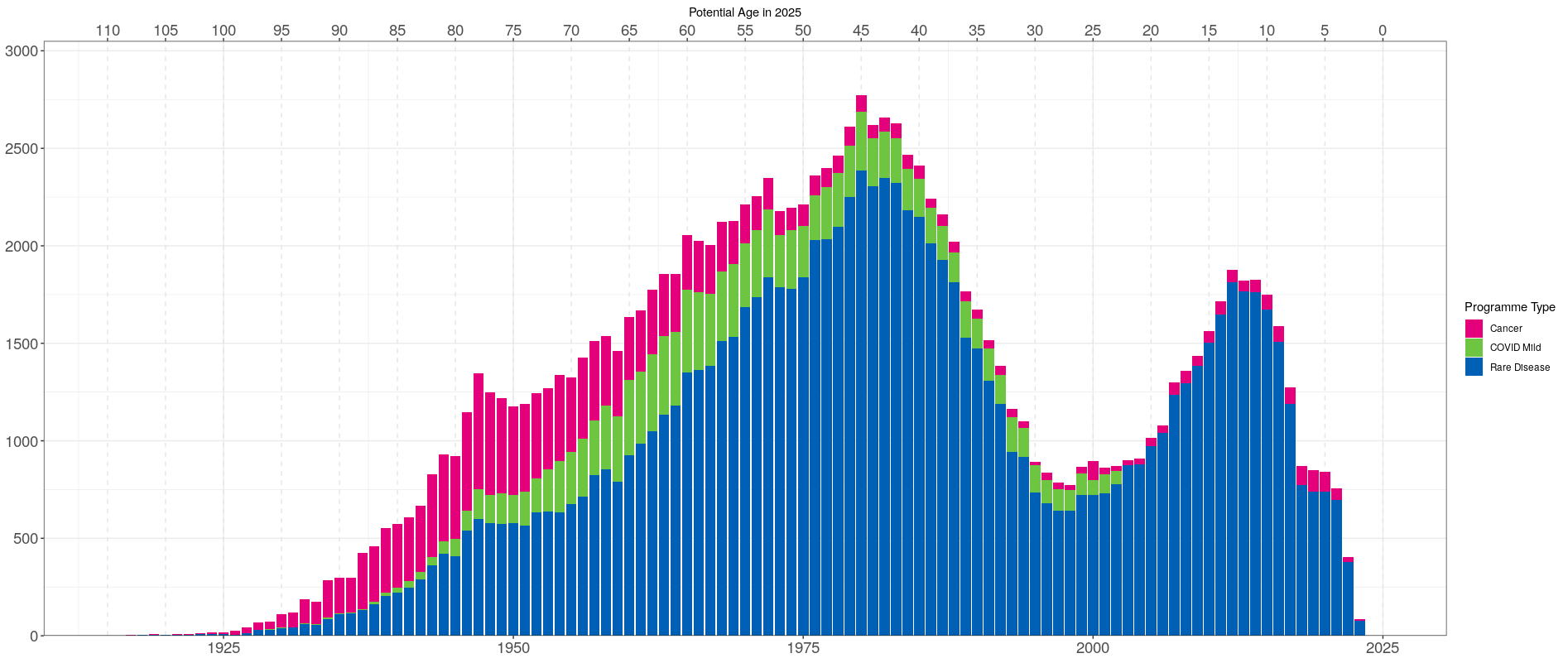
Below is a plot of the distribution of year of birth and estimated age in 2025 for individuals included in aggV3, grouped by programme of enrolment. Estimated age was calculated from each individual’s year of birth without accounting for potential death status. Individuals enrolled in the COVID Severe study have no available information on their year of birth and are therefore omitted from the plot (~7,000 participants).
Sex-chromosome ploidy was estimated as part of the DRAGEN 3.7.8 pipeline. Below is a breakdown of the percentage of individuals by estimated sex chromosome karyotype:
| Sex Chromosome Ploidy Estimation | Percentage of Samples |
|---|---|
| XX | 51.27% |
| XY | 48.32% |
| X0 | 0.27% |
| XXYY | 0.05% |
| XYY | 0.05% |
| XXX | 0.04% |
| XXXY | <5 samples |
We also provide a document containing bar charts that show the percentage of samples enrolled in each disease group or clinical indication across the 100kGP and NHS-GMS programmes. Some participants may have been enrolled under more than one disease group or clinical indication. In these cases, individuals may appear in multiple categories and be counted more than once in the charts.
Note on grouping of disease groups and clinical indications
For the above plots, all groups with 20 of fewer individuals were combined into aggregate groups. These are labeled Other disorders for rare disease-related plots and Other cancers for cancer-related plots.
The list below outlines the composition of each aggregate group:
- 100K Rare Disease: Infectious diseases
- 100K Cancer: NASOPHARYNGEAL,OTHER,SINONASAL
- NHS-GMS Rare disease: Holoprosencephaly - NOT chromosomal,Possible X-linked retinitis pigmentosa
- NHS-GMS Cancer: Acute Leukaemia Other,Adipocytic Soft Tissue Tumour Differential,ALK Negative Anaplastic Large Cell Lymphoma (Including Primary Cutaneous Subtypes),ALK Positive Anaplastic Large Cell Lymphoma,ALK Positive Large B Cell Lymphoma,Alveolar Soft Part Sarcoma,Anaplastic Astrocytoma - Paediatric,Angiomatoid Fibrous Histiocytoma,Aplastic Anaemia,Atypical Teratoid/Rhabdoid Tumour - Paediatric,B Cell Non-Hodgkin Lymphoma,Blastic Plasmacytoid Dendritic Cell Neoplasm,Bone Forming Bone Tumour Differential,Brain Tumour - No Further Morphological Classification - Paediatric,Burkitt Lymphoma,Cancer of Unknown Primary,Cartilage Forming Bone Tumour Differential,Chondroblastoma,Chondrosarcoma Conventional Central,Chronic Myeloid Leukaemia,Clear Cell Kidney Sarcoma - Paediatric,Clear Cell Sarcoma of Soft Tissue,CNS Ewing Sarcoma Family Tumour With CIC Alteration,Congenital Mesoblastic Nephroma - Paediatric,Cystic Nephroma - Paediatric,Dermatofibrosarcoma Protuberans,Desmoplastic Infantile Gangliogliomas - Paediatric,Desmoplastic Medulloblastoma - Paediatric,Desmoplastic Small Round Cell Tumour,Diffuse Astrocytoma - Paediatric,Diffuse Midline Glioma - Adult,Diffuse Midline Glioma - Paediatric,Embryonal Tumour Differential - Adult and Paediatric,Embryonal Tumours with Multi-Layered Rosettes - Paediatric,Ependymoma - Adult,Ependymoma - Paediatric,Epithelioid Soft Tissue Tumour Differential,Ewing Like Sarcoma/PNET,Ewing-Like Soft-Tissue Sarcoma,Extraskeletal Myxoid Chondrosarcoma,Giant Cell Tumour of Bone,Glial and Glioneuronal Tumour Differential - Paediatric,Glial Tumours - Paediatric,Glioblastoma - Paediatric,Glioma - Paediatric,High Grade Intrinsic Brain Tumour Differential - Adult,High Grade Lymphoma,High-Grade Neuroepithelial Tumour-Bcor Group,Histiocytosis,IDH-Wildtype Glioblastoma - Paediatric,Infantile Fibrosarcoma,Inflammatory Myofibroblastic Tumour,Juvenile Myelomonocytic Leukaemia,Low Grade Fibromyxoid Sarcoma,Low Grade Glioma - Adult,Low Grade Glioma - Paediatric,Low Grade Glioma/Glioneuronal Tumours - Adult,Low Grade Intrinsic Brain Tumour Differential - Adult,Lung - Paediatric,Lymphoma,MDS/MPN,Medulloblastoma - Paediatric,Medulloblastoma all Subtypes,Medulloblastoma Group 3/4 - Paediatric,Meningioma - Paediatric,Mesenchymal Chondrosarcoma,Myelodysplasia,Myeloproliferative Neoplasm,Myxoid Soft Tissue Tumour Differential,Myxoid/Round Cell Liposarcoma,Myxoinflammatory Fibroblastic Sarcoma,NK Cell/Gamma-Delta T Cell Lymphoma,Oligodendroglioma - Adult,Osteoclast-Rich Bone Tumour Differential,Ovarian Carcinoma,Phosphaturic Mesenchymal Tumour,Pilocytic Astrocytoma - Adult,Pineoblastoma - Paediatric,Pituitary Tumours,Pleomorphic Xanthoastrocytoma - Paediatric,Pleuropulmonary Blastoma - Paediatric,Primitive Mesenchymal Myxoid Tumour of Infancy,Pseudomyogenic Haemangioendothelioma,Radiation Induced Angiosarcoma,Rare Primitive Neuroectodermal Tumours Groups 2/3 - Paediatric,Renal Tumour Differential - Paediatric,Renal Tumours - Paediatric,Retinoblastoma - Paediatric,Rhabdoid Tumours - Paediatric,Rosette-Forming Glioneuronal Tumour - Paediatric,Round Cell Sarcoma Nos,Round Cell Sarcoma of Soft Tissue Differential,SHH Medulloblastoma - TP53 MUTANT - Paediatric,Spindle Cell Tumour of Bone Differential,Synovial Sarcoma,T Cell Non-Hodgkin Lymphoma,Testicular - Paediatric,Thyroid Papillary Carcinoma - Paediatric,Triple Negative Breast Cancer (WGS PILOT),Unable To Grade Intrinsic Brain Tumour - Adult,Undifferentiated Round Cell Sarcoma of Infancy,Uterine Sarcomas (Inc Endometrial),Vascular Soft Tissue Tumour Differential,Vascular Tumour of Bone Differential,Well Differentiated/Dedifferentiated Liposarcoma
Sample source and library preparation¶
The majority of samples are from blood. A small portion of samples are from various other sources outlioned below. Sample sources with fewer than five samples each were combined into a single OTHER category. This category includes samples taken from amniotic fluid and and chorionic villus sampling.
| Sample source | Percentage of samples |
|---|---|
| BLOOD | 98.8 |
| SALIVA | 0.597 |
| TISSUE | 0.249 |
| FRESH TISSUE IN CULTURE MEDIUM | 0.159 |
| BONE_MARROW | 0.111 |
| FIBROBLAST | 0.108 |
| GERMLINE | 0.02 |
| OTHER | 0.004 |
Most samples were prepared using EDTA, while the remaining proportion were prepared either fresh frozen or with Oragene.
| Sample Preparation method | Percentage of samples |
|---|---|
| EDTA | 98.9 |
| FF | 0.589 |
| ORAGENE | 0.549 |
Library preparation information was not available for a subset of 100kGP samples, resulting in partial missingness in the table below.
| Library preparation type | Percentage of samples |
|---|---|
| TruSeq PCR-Free High Throughput | 56.1 |
| NovaSeq TruSeq PCR-Free High Throughput | 37.3 |
| UNAVAILABLE | 6.56 |
| TruSeq Nano High Throughput | 0.052 |
| NovaSeq TruSeq PCR-Free High Throughput, Unknown | 0.012 |
| pcr | < 5 Samples |