Ensembl BioMart¶
Ensembl BioMart is a powerful web tool for performing complex querying and filtering of the various Ensembl databases (incl. Ensembl Genes, Ensembl Variation, and Ensembl Regulation). It is often used for ID mapping and feature extraction. Almost any data that is viewable in the Ensembl genome browser can be accessed systematically from BioMart. For more information about the tool and its utility, please the see external Ensembl documentation outside of the Research Environment.
Inside the Research Environment, you may access Ensembl BioMart and download publicly accessible information in two ways:
- Via a desktop shortcut to the BioMart URL
- Via the biomaRt package
Accessing Ensembl BioMart via URL¶
To access Ensembl BioMart via the web, you may visit the following whitelisted URL in your browser: https://www.ensembl.org/biomart/martview
From the desktop, you may simply click the 'Ensembl BioMart' icon:

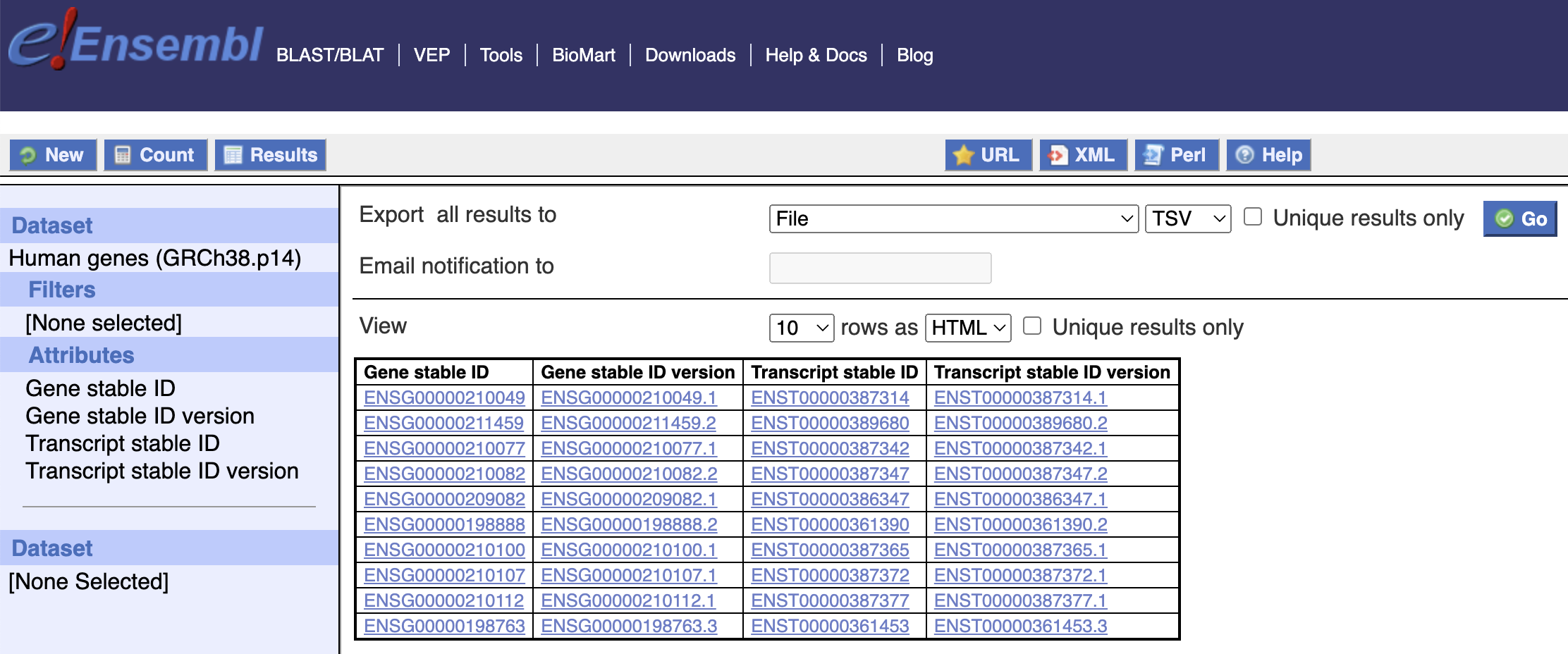
Once launched, you may select your dataset of interest, choose bespoke filters and attributes, and export results as TSV directly to your Research Environment session.
Accessing Ensembl BioMart via the biomaRt package¶
Accessing biomaRt from pre-installed locations¶
To access Ensembl BioMart programmatically in an R session, you may use the biomaRt R package.
In the Research Environment biomaRt is pre-installed for R 4.0.2 and 4.2.1 in the following locations:
/tools/aws-workspace-apps/ce/R/4.0.2/tools/aws-workspace-apps/ce/R/4.2.1
biomaRt is pre-installed for R version 4.2.1 on the HPC without accessing the above library locations.
Self-installation of biomaRt¶
After you have set up your .Rprofile, .Renviron, and .netrc configuration files, you may also install biomaRt into your own R package library location using BioConductor:
if (!require("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("biomaRt")
To load the package:
Using biomaRt - External resources¶
For the best documentation on using biomaRt to its full potential, please visit the external documentation from Bioconductor and Ensembl.