100kGP General clinical data¶
A number of tables in LabKey are common to all participants, both cancer and rare disease participants. These cover data about the participants themselves, the samples, results of bioinformatic analyses and participant medical history. All tables and their fields are described in our data dictionary.
Primary and secondary data tables
Primary clinical data were collected when participants were enrolled in the programme; tables are tagged with .
Secondary clinical data were obtained from third parties such as NHSE; tables are tagged with .
Participant information tables¶
| LabKey table | Description | Primary or secondary | CloudOS tsv filename |
|---|---|---|---|
participant_summary |
an aggregation of data from several Rare Diseases, Cancer and Bioinformatics tables within LabKey which aims to provide commonly used fields across all these tables in a single space for all our participants. The Participant Summary table is composed of demographic information from table participant, ancestry information from aggregate_gvcf_sample_stats, death details from tables death_details, mortality and rare_diseases_pedigree_member, study information from cancer_analysis, disease and phenotype information from tables rare_diseases_participant_disease, rare_disease_interpreted rare_diseases_participant_phenotype (filtered for phenotypes where hpo_present = 'yes') and cancer information from tables cancer_staging_consolidated and cancer_participant_disease. This table was previously named key_columns in v16. |
||
participant |
contains personal information (such as relatives or self-reported ethnicity), points of contact with the Project (e.g. handling Genomic Medicine Centre or Trust), and a record of the status of their clinical review. | gel_participant_100k.tsv |
|
death_details |
contains participant deaths submitted by Genomic Laboratory Hubs, likely less complete than the data collected by ONS and NHSE. | gel_gmc_death_details_100k.tsv |
participant_summary details
The participant_summary table was generated using the following tables/calculations:
| field | origin table(s) |
|---|---|
participant_stated_gender |
participant |
participant_karyotyped_sex |
rare_disease_analysis, cancer_analysis |
participant_phenotyped_sex |
participant |
yob |
participant |
genotyped_ancestry |
aggregate_gvcf_sample_stats (filtered on pred_ancestry value >= 0.8) |
predominant_ancestry |
aggregate_gvcf_sample_stats |
predominant_ancestry_value |
aggregate_gvcf_sample_stats |
programme |
participant |
participant_type |
participant |
affection_status |
rare_disease_interpreted |
date_of_consent |
participant |
cancer_diagnosis_date |
cancer_staging_consolidated (calculated MINDATE(diagnosis_date) group by participant_id) |
cancer_diagnosis_age |
calculated (DATEDIF(yob, cancer_diagnosis_date, year)) |
cancer_study_name |
cancer_analysis |
cancer_study_abbreviation |
cancer_analysis |
cancer disease_type |
cancer_participant_disease |
cancer_disease_sub_type |
cancer_participant_disease |
disease_diagnosis_date |
rare_diseases_participant_disease |
disease_diagnosis_age |
calculated (DATEDIF(yob, disease_diagnosis_date, year))` |
normalised_disease_group |
rare_diseases_participant_disease |
normalised_disease_sub_group |
rare_diseases_participant_disease |
normalised_specific_disease |
rare_diseases_participant_disease |
hpo_term |
rare_diseases_participant_phenotype (filtered for participants where hpo_present = 'yes') |
death_date |
death_details, mortality,rare_diseases_pedigree_member` |
Sampling and sequencing tables¶
| LabKey table | Description | Primary or secondary | CloudOS tsv filename |
|---|---|---|---|
clinic_sample |
describes the taking and handling of participant samples at the Genomic Medicine Centres, i.e. in the clinic, as well as the type of samples obtained. Because of the complexities of handling and managing tumour tissues samples in a clinical setting, there are many fields that are cancer-specific. | gel_clinic_sample_100k.tsv |
|
clinic_sample_quality_check_result |
describes the quality control of obtaining and handling participant samples at the Genomic Medicine Centres, i.e. in the clinic. | gel_clinic_sample_qc_results_100k.tsv |
|
laboratory_sample |
describes the handling of samples at the biorepository and in preparation for sequencing, as well as the type of biological sample. | gel_laboratory_sample_100k.tsv |
|
laboratory_sample_omics_availability |
contains information on other biological samples that are available in our biobank for our participants as of the latest data release. Please note that these samples have not been sequenced nor analysed by Genomics England. | gel_laboratory_sample_omics_100k.tsv |
|
plated_sample |
contains, for each sequenced sample, the plate key and plate id, along with Illumina QC date, status and few other QC information. | gel_plated_sample_100k.tsv |
Results of bioinformatic analysis¶
These tables contain data from and information about Genomics England interpretation pipelines for participants from both cancer and rare disease programmes. These tables do not directly include primary and secondary sources of clinical data.
| LabKey table | Description | Primary or secondary | CloudOS tsv filename |
|---|---|---|---|
aggregate_gvcf_sample_stats |
contains the samples that have been used to create the aggregate vcf files (/gel_data_resources/main_programme/aggregated_illumina_gvcf/GRCH38/20190228/) and their QC metrics. Individual sample QC data was retrieved from Genomics England OpenCGA database. Most sequencing metrics are BAM file statistics provided from Illumina or Genomics England WGS data processing pipeline. The table contains principal components, a set of unrelated individuals and probabilities of ancestry membership, and more. |
gel_rare_disease_and_germline_genomic_variant_call_format_sample_statistics_100k.tsv |
|
genome_file_paths_and_types |
contains the folder location for the bam and vcf files for each participant. Please be aware that the same genome can be released with multiple versions of mapping/variant calling pipeline. Since the Main Programme Data Release Version 10, we have added file paths to genomes that have been realigned with the DRAGEN pipeline. Filter for 'DRAGEN_Pipeline2.0' in the 'Delivery Version' column to access them. |
||
sequencing_report |
contains data describing the sequencing of their genome(s) and associated output, as well as the sample type that the sequence is from, e.g. rare disease germline, cancer somatic, etc. | gel_sequencing_report_100k.tsv |
|
panels_applied |
contains the name and version of the panel(s) that was applied to their genome. | gel_panels_applied_100k.tsv |
Long Read Sequencing¶
Contains tables related to long-reads sequencing data for 100,000 Genomes Project participants.
| LabKey table | Description | Primary or secondary | CloudOS tsv filename |
|---|---|---|---|
lrs_laboratory_sample |
Data describing the characteristics and processing methods (DNA to library preparation) of samples from participants in the 100,000 Genomes Project for which long-reads sequencing has been carried out. | gel_lrs_laboratory_sample_100k.tsv |
|
lrs_sequencing_data |
This table includes data describing long-read sequencing of a subset of 100,000 Genomes Project participants and associated output, including paths to raw and BAM files. | gel_lrs_sequencing_data_100k.tsv |
|
cancer_ont_cohorts |
Table listing participant ids, sample data, file paths and sequencing statistics for Oxford Nanopore cancer cohorts available in the Research Environment, along with corresponding matched germline and Illumina short reads files where available | ||
rare_disease_pacbio_pilot |
This is a dataset of 91 rare disease samples from the 100kGP genome project re-sequenced with Pacific Biosciences (PacBio) as an example dataset to to demonstrate the utility of their HiFi technology. |
Participant medical history¶
Secondary Clinical Data is available for 100kGP participants with current consent. Clinical data is not available for participants that have withdrawn from the 100kGP or were otherwise ineligible. Up to Data Release V14 any participant on child consent who turned 16 without reconsenting as an adult was deemed ineligible and removed from the release. For Data Release V14 a decision was made to reinclude these participants with a caveat that where a participant was consented as a child but turned 16 without re-consenting as an adult, only data collected under that consent (i.e. all that collected prior to their 16th birthday) is released.
Some participants' secondary data records may have a sudden end point that does not correlate to an end to treatment. The column secondary_data_received_until_year in the table participant contains the year that the participant turned 16 or this column will be null.
Hospital Episodes Statistics from NHSE¶
Hospital Episodes Statistics (HES) contain details of all admissions, outpatient appointments, critical care and A&E attendances at NHS hospitals in England. Each data entry is collected during a patient's time in hospital and are submitted to allow hospitals to be paid for the care they deliver. HES data are designed to enable secondary use, that is use for non-clinical purposes, of these administrative data.
It is a records-based system that covers all NHS trusts in England, including acute hospitals, primary care trusts and mental health trusts. HES information is stored as a large collection of separate records and Genomics England receives regular partial exports of HES data held for each of the participants within the 100,000 Genomes Project, which are linked with their Participant ID. HES data are presented in LabKey as separate datasets.
The HES data are presented in LabKey with each row representing a separate period of care for that participant. Therefore, each participant may have one or more rows of data. Often there will be empty fields, due to the way the data is structured.
| LabKey table | Description | Primary or secondary | CloudOS tsv filename |
|---|---|---|---|
hes_ae |
accident and emergency; contains historic records of A&E attendances | nhs_d_hospital_episodes_statistics_accident_and_emergency_100k.tsv |
|
hes_apc |
admitted patient care; contains historic records of admissions into secondary care. | nhs_d_hospital_episodes_statistics_admitted_patient_care_100k.tsv |
|
hes_cc |
critical care; contains historic records of admissions into critical care. | nhs_d_hospital_episodes_statistics_critical_care_100k.tsv |
|
hes_op |
outpatient; contains historic records of outpatient attendances. | nhs_d_hospital_episodes_statistics_outpatient_100k.tsv |
|
ecds |
Main dataset of urgent and emergency care. Expands hes_ae and will replace it entirely in the future. | nhs_d_emergency_care_dataset_100k.tsv |
Some data-points, such as diagnoses and treatments, are split across multiple columns since there will be multiple entries per visit. There are also columns that concatenate these values together, making them easier to search.
Concatenated columns available
The concatenated columns in each of the tables are shown in the table below:
| Table | Concatenated Column Name | Source Columns |
|---|---|---|
ecds |
care_professional_tier_all |
care_professional_tier_01 - care_proffessional_tier_10 |
ecds |
classification_all |
classification_01 - classification_04 |
ecds |
comorbidities_all |
comorbidities_01 - comorbidities_10 |
ecds |
diagnosis_code_all |
diagnosis_code_01 - diagnosis_code_12 |
ecds |
diagnosis_qualifier_all |
diagnosis_qualifier_01 - diagnosis_qualifier_12 |
ecds |
drug_alcohol_code_all |
drug_alcohol_code_01 - drug_alcohol_code_04 |
ecds |
investigation_code_all |
investigation_code_01 - investigation_code_12 |
ecds |
treatment_code_all |
treatment_code_01 - treatment_code_12 |
hes_apc |
acpdisp_all |
acpdisp_1 - acpdisp_9 |
hes_apc |
acpdqind_all |
acpdqind_1 - acpdqind_9 |
hes_apc |
acploc_all |
acploc_1 - acploc_9 |
hes_apc |
acpout_all |
acpout_1 - acpout_9 |
hes_apc |
acpsour_all |
acpsour_1 - acpsour_9 |
hes_apc |
acpspef_all |
acpspef_1 - acpspef_9 |
hes_apc |
diag_all |
diag_01 - diag_20 |
hes_apc |
opertn_all |
opertn_01 - opertn_24 |
hes_ae |
diag_all |
diag_01 - diag_12 |
hes_ae |
diag2_all |
diag2_01 - diag2_12 |
hes_ae |
diaga_all |
diaga_01 - diaga_12 |
hes_ae |
diags_all |
diags_01 - diags_12 |
hes_ae |
invest_all |
invest_01 - invest_12 |
hes_ae |
invest2_all |
invest2_01 - invest2_12 |
hes_ae |
treat2_all |
treat2_01 - treat2_12 |
hes_ae |
treat_all |
treat_01 - treat_12 |
hes_op |
diag_all |
diag_01 - diag_12 |
hes_op |
opertn_all |
opertn_01 - opertn_24 |
mortality |
icd10_multiple_cause_all |
icd10_multiple_cause_01 - icd10_multiple_cause_15 |
Diagnosis and treatment codes
ICD-10¶
ICD-10 is a classification of diseases that allows systematic recording, analysis, interpretation and comparison of mortality and morbidity data. It is the international standard diagnostic classification for all general epidemiological and many health-management purposes. Although the ICD is primarily designed for the classification of diseases and injuries with a formal diagnosis, not every problem or reason for coming into contact with health services can be categorised in this way.
ICD-10 codes must be used in the manner set forth in Volume 2: | | Instruction Manual of the International Statistical Classification of Diseases and Related Health Problems, Tenth Revision. You are responsible for ensuring that the codes are properly used in this manner.
For more information on ICD-10, please see the 'International statistical classification of diseases and related health problems (ICD-10)' document.
ICD codes and code descriptions are deposited in the Research Environment under the folder: | | /gel_data_resources/licenced_resources/ICD10
ICD-O-3¶
The International Classification of Diseases for Oncology (ICD-O) is internationally recognised as the definitive classification of neoplasms. It is used by cancer registries throughout the world to record incidence of malignancy and survival rates, and the data produced are used to inform cancer control, research activity, treatment planning and health economics. The classification of neoplasms used in ICD-O links closely to the definitions of neoplasms used in the WHO/IARC Classification of Tumours series, which are compiled by consensus groups of intenational experts and, as such, the classification is underpinned by the highest level of scientific evidence and opinion.
ICD-O consists of two axes (or coding systems), which together describe the tumour:
- the
topographicalcode, which describes the anatomical site of origin (or organ system) of the tumour - the
morphologicalcode, which describes the cell type (or histology) of the tumour, together with the behaviour (malignant or benign).
SNOMED¶
SNOMED was started in 1965 as a Systematised Nomenclature of Pathology (SNOP) and was further developed into a logic-based health care terminology. SNOMED CT was created in 1999 by the merger, expansion and restructuring of two large-scale terminologies: SNOMED Reference Terminology (SNOMED RT) and the Clinical Terms Version 3 (CTV3) (formerly known as the Read codes), developed by the NHS.
Other medical history tables¶
| LabKey table | Description | Primary or secondary | CloudOS tsv filename |
|---|---|---|---|
did |
contains historic diagnostic imaging records. | nhs_d_diagnostic_imaging_metadata_100k.tsv |
|
mortality |
lists the Office of National Statistics' cause of death records. | office_of_national_statistics_mortality_100k.tsv |
Mental health data¶
Mental Health Datasets contain historic data on patients receiving care in NHS specialist mental health services. Note that mental health (MH) data is split into mental health minimum data (mhmd_), mental health learning disabilities dataset (mhldds_) and mental health services dataset (mhsds_).
mhmd/_*(Mental Health Minimum Dataset) - 2011-2014mhldds/_*(Mental Health Learning Disabilities Dataset) - 2014-2016mhsds/_*(Mental Health Services Dataset) - 2016 onwards
MHMD and MHLDDS each consist of three tables; event, record and episode, covering the periods 2011-2014 and 2014-2016 respectively. MHSDS is based on a new data model and consists of 35 tables covering the period 2016-2019. Thus, events that happened until March 2014 are found in mhmd_*, between April 2014 and April 2016 in mhldds_* and after April 2016 in mhsds_*.
To make the new MHSDS dataset more accessible we have also generated four curated overview tables as well as a flag table showing which participants have data in individual MHSDS tables.
mhsds tables¶
| LabKey table | Description | Primary or secondary | CloudOS tsv filename |
|---|---|---|---|
mhsds_master_patient_index |
Provides patient information, demographics and death details for participants present in the MHSDS dataset. One record per participant per recording period (2016/17, 2017/18, 2018/19) giving a maximum of three records per participant. | ||
mhsds_gp_practice_registration |
Carries details of GP Practice registrations for participants present in the MHSDS dataset. One record per change of GP practice registration. | ||
mhsds_patient_indicators |
Carries details of specific indicators relating to the patient including psychosis. | ||
mhsds_care_coordinator |
Carries details of the mental health care coordinator assigned to a patient. One record per assignment. | ||
mhsds_care_plan_type |
Carries details of Care Plans created for a patient by their responsible organisation. One record per care plan created for the patient. | ||
mhsds_crisis_plan |
The predecessor (pre 2018) to Care Plan Type. Carries detail of Crisis Plans created for a patient by their responsible organisation. One record per crisis plan created for the patient. Other care plan types are not included. | ||
mhsds_care_plan_agreement |
Carries detail of any agreements to Care Plans by a patient, team or organisation. One record per care plan agreement. | ||
mhsds_service_or_team_referral |
Carries details of the referral that a patient is subject to. This is an instance where a patient is referred to specialist care. One record for each referral. | ||
mhsds_service_or_team_type_referred_to |
Carries details of the service or team that a patient is referred to. One record for each service or team referred to. | ||
mhsds_other_reason_for_referral |
Carries additional details about why a patient has been referred to a specific service. One record per additional referral. | ||
mhsds_referral_to_treatment |
Carries details about the referral to treatment details for the patients referral. One record for each Referral To Treatment period. | ||
mhsds_onward_referral |
Carries details about any onward referral of a patient. One record per onward referral. | ||
mhsds_discharge_plan_agreement |
Carries details about any agreements to a Discharge Plan by a person, team or organisation. One record per agreement of a discharge plan. | ||
mhsds_care_contact |
Carries details about any contacts with a patient which have taken place as part of a referral. One record per care contact. | ||
mhsds_care_activity |
Carries details about any Care Activity undertaken at a Care Contact. One record per care activity. | ||
mhsds_other_in_attendance |
Carries details about any other people in attendance at a Care Contact. One record per other person in attendance at a Care Contact. | ||
mhsds_indirect_activity |
Carries details of indirect activity which takes place as a result of the referral. One record for each instance of indirect activity taking place. | ||
mhsds_responsible_clinician_assignment |
Carries details of the assignment of a Mental Health Responsible Clinician to the patient. One record per assigned Mental Health Responsible Clinician. | ||
mhsds_hospital_provider_spell |
Carries details of each Hospital Provider Spell for a patient. One record per hospital provider spell. This is a continuous period of inpatient care under a single Hospital Provider starting with a hospital admission and ending with a discharge from hospital. | ||
mhsds_ward_stay |
Carries additional details of Ward Stays which occurred during a Hospital Provider Spell for the patient. One record per ward stay. | ||
mhsds_assigned_care_professional |
Carries details of the Care Professional assigned responsibility for the care of the patient. One record per care professional admitted care episode. | ||
mhsds_delayed_discharge |
Carries details of the patient's Mental Health Delayed Discharge Periods which occurred during a Hospital Provider Spell. One record per instance of a patient being subject to a mental health delayed discharge period. | ||
mhsds_hospital_provider_spell_commissioner |
Carries details of each Commissioner Assignment Period during a Hospital Provider Spell. One record per commissioner assignment period. | ||
mhsds_medical_history_previous_diagnosis |
Carries details any previous diagnoses for a patient which are stated by the patient or recorded in medical notes. These do not necessarily have to have been diagnosed by the organisation submitting the data. One record per previous diagnosis. | ||
mhsds_provisional_diagnosis |
Carries details of a provisional diagnosis recorded for a patient. One record per provisional diagnosis. | ||
mhsds_primary_diagnosis |
Carries details of the primary diagnosis recorded for the patient. Only one record is permitted for the primary diagnosis per patient. | ||
mhsds_secondary_diagnosis |
Carries details of a secondary diagnosis recorded for a patient. One record for each secondary diagnosis. | ||
mhsds_coded_scored_assessment_referral |
Carries details of scored assessments that are issued and completed as part of a referral to a mental health service, but do not take place at a specific contact. One record per coded scored assessment question or dimension captured outside of a Care Contact. | ||
mhsds_coded_scored_assessment_act |
Carries details of scored assessments that are issued and completed as part of a specific Care Activity. One record per coded scored assessment question or dimension captured as part of a specific Care Activity. | ||
mhsds_coded_scored_assessment_cont |
Replaced by coded_score_assessment_act in 2018. Carries details of scored assessments that are issued and completed as part of a specific Care Activity. One record per coded scored assessment question or dimension captured as part of a specific Care Activity. | ||
mhsds_care_programme_approach_care_episode |
Carries details of the periods of time the patient spent on Care Programme Approach. One record per CPA Care Episode. | ||
mhsds_care_programme_approach_review |
Carries details of the Care Programme Approach (CPA) reviews undertaken for the patient. One record permitted for the most recent CPA Review that has taken place. | ||
mhsds_clustering_tool_assessment |
Carries details of all clustering tool assessments for all patients. One record per Clustering Tool Assessment. | ||
mhsds_coded_score_assessment_clustering_tool |
Carries details of scored assessments that are issued and completed as part of a Clustering Tool assessment. One record per coded scored assessment question or dimension captured as part of a Clustering Tool assessment. | ||
mhsds_care_cluster |
Carries details of the Care Cluster resulting from a clustering tool assessment. One record per period of time that a patient was allocated to a Care Cluster. |
The mhsds curated tables have been generated by joining the main columns of interest from the individual MHSDS tables into the following four overview tables:
| LabKey table | Description | Primary or secondary | CloudOS tsv filename |
|---|---|---|---|
mhsds_curated_participant |
Overview of general participant information; demographics, death details, GP registrations and psychosis indicators, as well as details of any care plans created for a participant; 2016 onwards. | ||
mhsds_curated_community |
Overview of community (outpatient) care. This includes details on referrals, discharge agreements and care contacts with associated care activities; 2016 onwards. | ||
mhsds_curated_inpatient |
Overview of inpatient care. This includes details of hospital spells, ward stays, delayed discharge periods and associated clinicians and care professionals; 2016 onwards. | ||
mhsds_curated_assessment_diagnoses_and_cluster |
Overview of scored assessments and clustering tool assessments completed, patient diagnoses and allocated care clusters; 2016 onwards. |
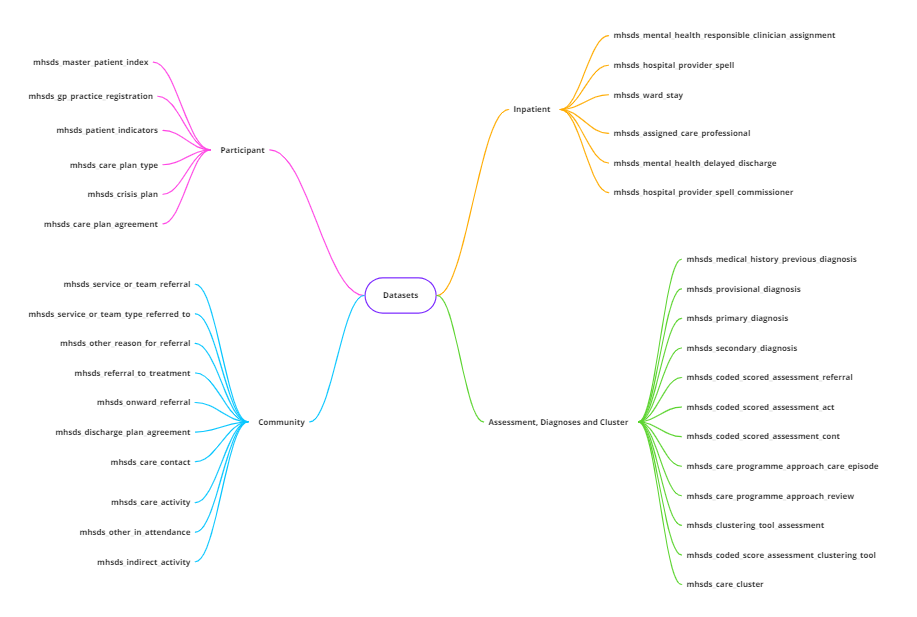
These curated tables give an overview of the data available across the entire dataset, giving a subset of the main data. Some rows have been removed from the curated tables due to data quality issues. For master_patient_index results in the Participant curated table, only the most recent data for each participant has been included.
Where column names are shared between datasets in the curated tables(e.g. codeproc), the column names have been prefixed with the initials of their dataset name - e.g. those with shared column names from mhsds_indirect_activity have been prefixed with ia_.
In the data dictionary, you will see the column 'Column Origin (Curated only)', which links together the curated columns in the format
An additional flag table has also been generated to provide an easy way to view which participants are present in which of the individual tables.
| LabKey table | Description | Primary or secondary | CloudOS tsv filename |
|---|---|---|---|
mhsds_dataset_flags |
contains, for each participant in mhsds, a flag of which mhsds tables they appear in. This only applies to the individual tables, not the curated tables. |
Each row represents a participant with true/false Boolean columns for each of the 35 mhsds tables. If a participant is not present in this table then they have no data available in the MHSDS dataset.
mhmd (2011-2014) and mhldds (2014-2016) individual tables¶
| LabKey table | Description | Primary or secondary | CloudOS tsv filename |
|---|---|---|---|
mhldds_episode |
contains historic records of mental health (MH) related admissions of Genomics England main programme participants. Episode and event tables link to the records table via mhm_mhmds_spell_id. | nhs_d_mental_health_learning_and_disability_data_set_episodes_100k.tsv |
|
mhldds_event |
contains historic records of MH related admissions of Genomics England main programme participants. Episode and event table link to the records table via mhm_mhmds_spell_id. | nhs_d_mental_health_learning_and_disability_data_set_events_100k.tsv |
|
mhldds_record |
contains historic records of MH related admissions of Genomics England main programme participants. One record per spell per patient in a provider. | nhs_d_mental_health_learning_and_disability_data_set_records_100k.tsv |
|
mhmd_v4_episode |
contains historic records of MH related admissions of Genomics England main programme participants. Episode and event tables link to the records table via mhm_mhmds_spell_id. | nhs_d_mental_health_minimum_dataset_episodes_100k.tsv |
|
mhmd_v4_event |
contains historic records of MH related admissions of Genomics England main programme participants. Episode and event tables link to the records table via mhm_mhmds_spell_id. | nhs_d_mental_health_minimum_dataset_events_100k.tsv |
|
mhmd_v4_record |
contains historic records of MH related admissions of Genomics England main programme participants. One record per spell per patient in a provider. | nhs_d_mental_health_minimum_dataset_records_100k.tsv |
Secondary Data - COVID¶
This data describes extracts from the NHSE microbiology results database, known as SGSS, following linkage to external cohort studies. Linkage is made by NHS number, and all positive and negative results are included.
Please be aware that:
- NHSE is integrating data from a large number of NHS laboratories and third party organisations, at a very rapid rate.
- Not all laboratories are reporting negative results.
- It is possible that duplicate entries may exist, because some laboratories' results may reach SGSS via several different routes.
- Results from NHS providers are all integrated at present.
- Data from the Milton Keynes Superlab and similar Academic/Industry partnerships have now been integrated. These initiatives are at present being used predominantly to test Health Care Workers. 'Healthcare Worker Testing' is a category in SGSS that's included in the result set. It's also worth noting that this automatically clears the inpatient indicator (as this category of tests are for hospital staff, not patients).
- As of 16th March 2020, when the UK entered the 'delay' phase of the outbreak, testing was largely restricted to those referred to hospital, who are likely to be on the severe end of the disease spectrum. Admission to hospital for infection control reasons alone has not been practiced in the delay phase. Therefore, positive results from those for whom there is evidence (from the microbiology record) of hospitalisation are likely to be derived from cases of clinically significant COVID disease. All tests with a sample date earlier than 16th March 2020 are excluded from the returned results.
- The SGSS database does not contain clinical information.
- The date of death is obtained by the Office for National Statistics linkage on the date of the extract.
- When we extract weekly data, we will re-extract all records linking to the NHS number list in the external cohort.